
AI agents build AI biology models
AI foundation models are rapidly reshaping biology, enabling tasks from protein structure prediction to cellular simulation. However, adapting these models to new datasets or experimental contexts remains technically demanding, limiting their use to a small set of experts. This creates a bottleneck in translating model advances into practical scientific workflows. Biomni Lab addresses this by enabling natural language-driven design, pre-training, and fine-tuning of biological models. By automating the full modeling lifecycle, it makes advanced AI methods accessible to a broader range of scientists. We’re looking for beta testers. Signup here for early access: https://forms.gle/1yhCP6Vrc12DaS4q6
The past two years have seen a wave of AI foundation models reshaping biology. Boltz predicts target protein structure while its companion BoltzGen generates binders for it, AlphaGenome maps sequence to expression, and virtual cell models like STATE simulate perturbations. Bessemer recently noted that nearly 400 models were released last year. These approaches are a new tool to help biologists with drug discovery and development and are increasingly the first step before beginning work in the wet lab.
A gap remains in the accessibility of these models; one that is not strictly limited to being able to run the models or set parameters. Scientists across academia and pharma want to adapt these models by finetuning them on proprietary datasets, adapting them to new assays, or pre-training versions tailored to specific species or cell types. But doing this today requires a rare combination of skills: deep learning, biology, and distributed systems. Today, only a relatively small number of people can execute this end-to-end.
AI can help.
We’re introducing a preview feature in Biomni Lab that lets any scientist design, pre-train, and fine-tune foundation models using natural language. You describe your data, your objective, and your compute budget, and the agent handles the rest—from literature review and experiment design to code generation, GPU orchestration, evaluation, and iteration—ultimately delivering a deployed model.
Biomni Lab operates across three modes that share a common agent loop. In Finetune, the agent adapts existing models using the appropriate strategy (full fine tuning, LoRA, or adapters), prepares the data, tunes hyperparameters, and evaluates performance. In Pre-train, it builds models from scratch, handling architecture design, distributed training, and optimization across GPU clusters. In Design, the agent starts from a task description, surveys existing approaches, proposes candidate architectures, implements them, and iterates—including creating entirely new model designs when needed.
Demonstrations
We validated this new capability across a range of real biological tasks, each executed end-to-end from a natural language prompt.
Biomni first took on regulatory genomics: fine-tuning Borzoi on 365K lentiMPRA regulatory elements across three cell types (K562, HepG2, WTC11). The fine-tuned model reached Pearson r=0.837 overall — a +0.285 gain over random initialization.
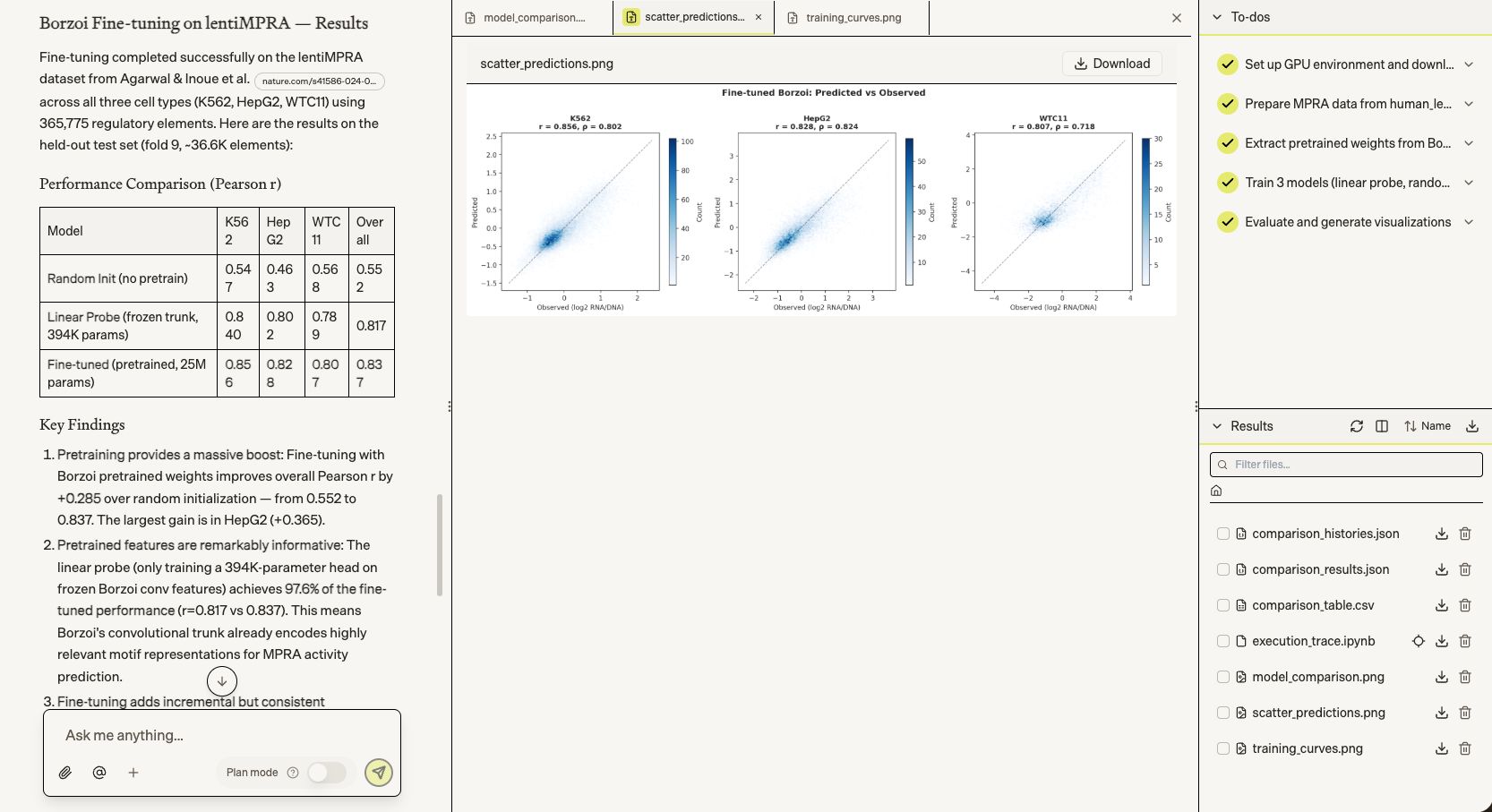
Biomni then tackled single-cell perturbation outcome prediction — finetuning scGPT for the Arc Virtual Cell Challenge on H1 hESC CRISPRi data. In 19 minutes, it took a zero-shot model with negative predictive correlation (DES=-0.75) to one that correctly captures perturbation direction across all 100 test conditions (DES=+0.41), with MAE dropping 40× from 29.6 to 0.72.
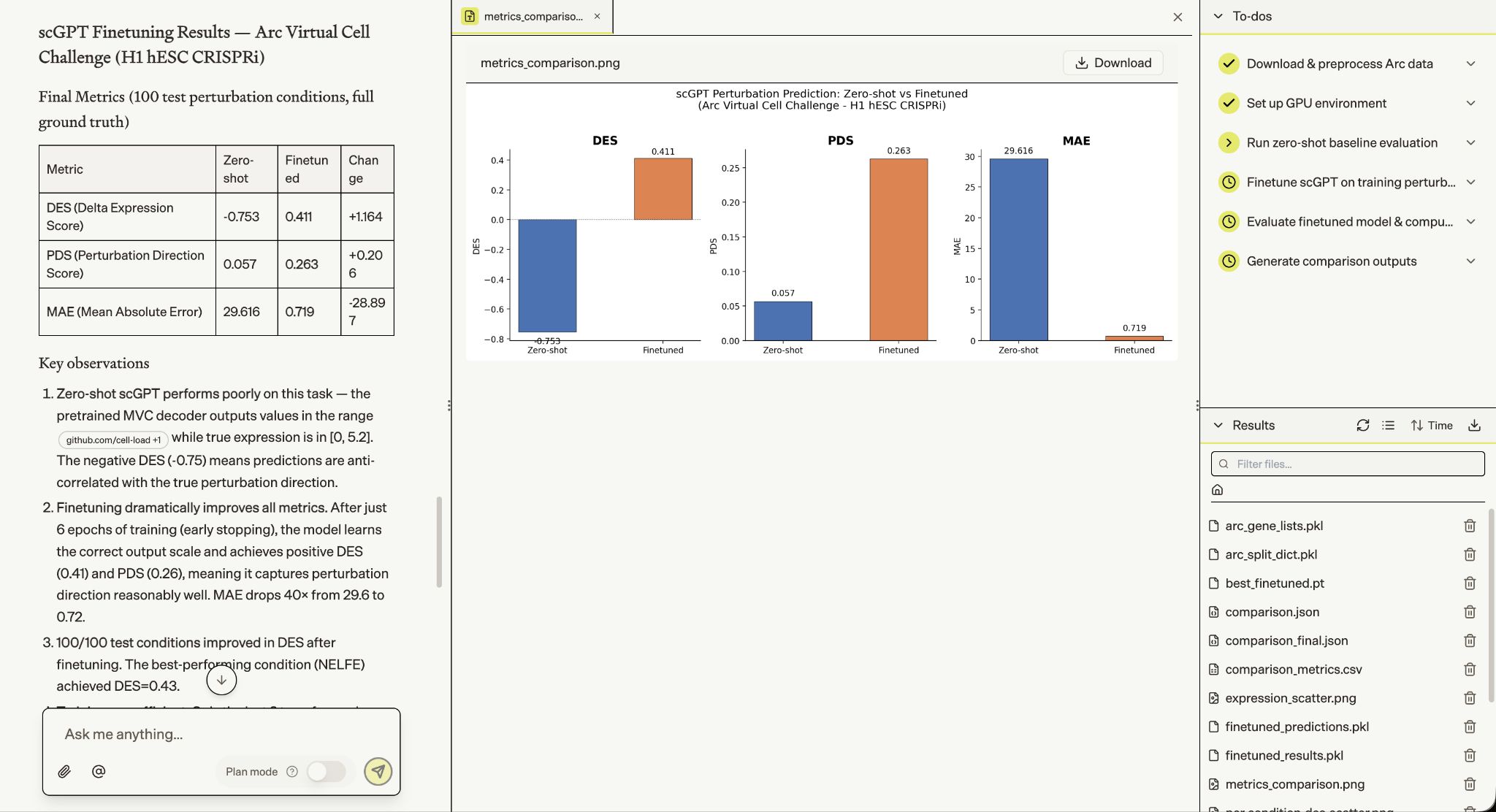
Biomni then fine-tuned ESM2-8M on subcellular localization prediction (DeepLoc-1.0), taking the model from 12% to 71.7% accuracy across 10 compartment classes in 5 epochs. On well-represented classes — Extracellular, Plastid, Nucleus, Mitochondrion — it hit F1 scores of 0.79–0.88, with failures concentrated in three rare classes comprising under 3% of training data.
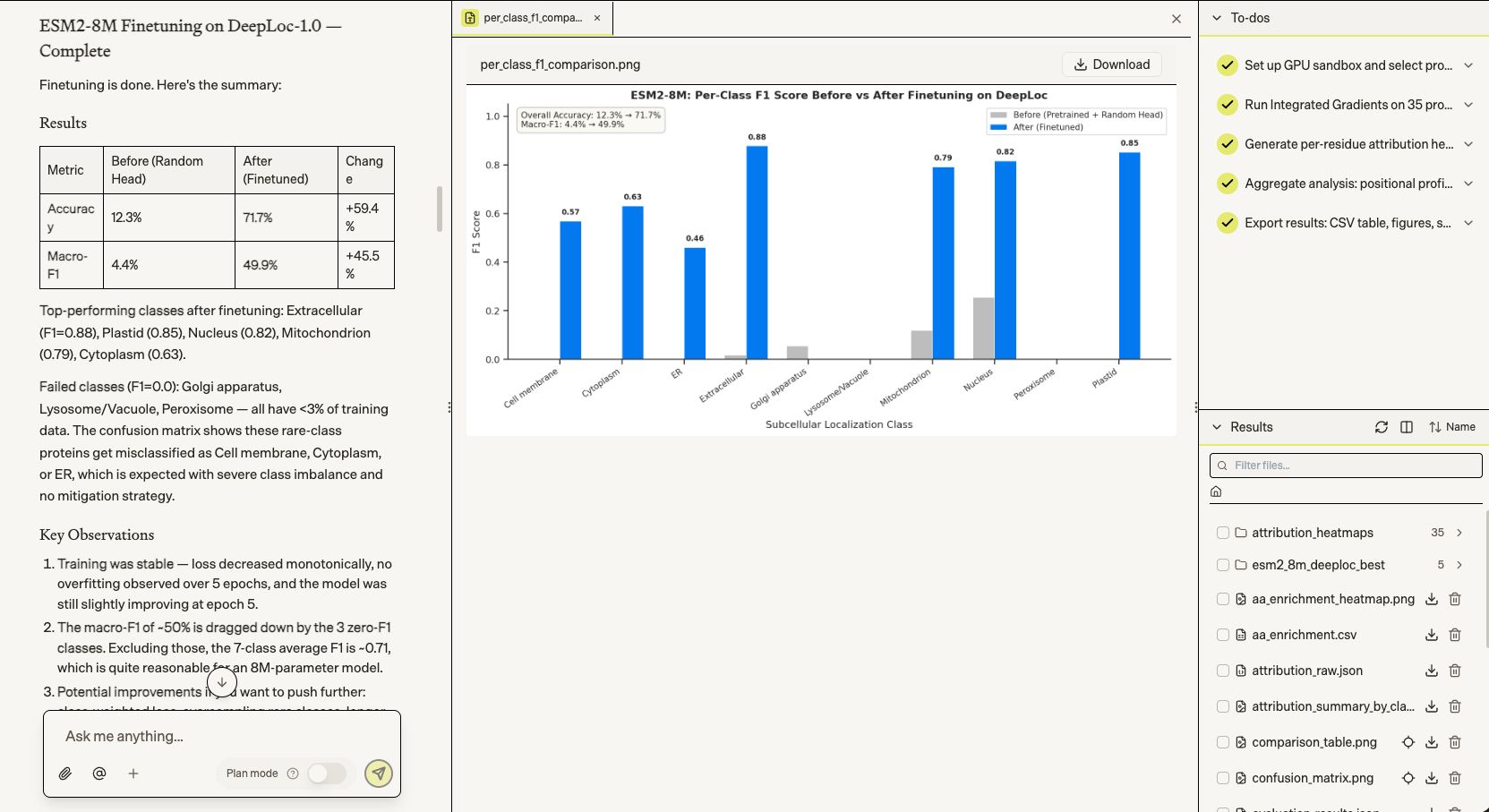
Biomni didn't stop at accuracy numbers — we asked it in plain language to explain why the model makes its predictions. It ran Integrated Gradients attribution analysis across 35 proteins and surfaced biologically coherent findings: N-terminal signal peptides driving Plastid and Extracellular predictions, K/R enrichment at NLS motifs for Nucleus, transmembrane domain signatures for Cell membrane. A single natural language prompt turned a trained model into a mechanistic hypothesis.
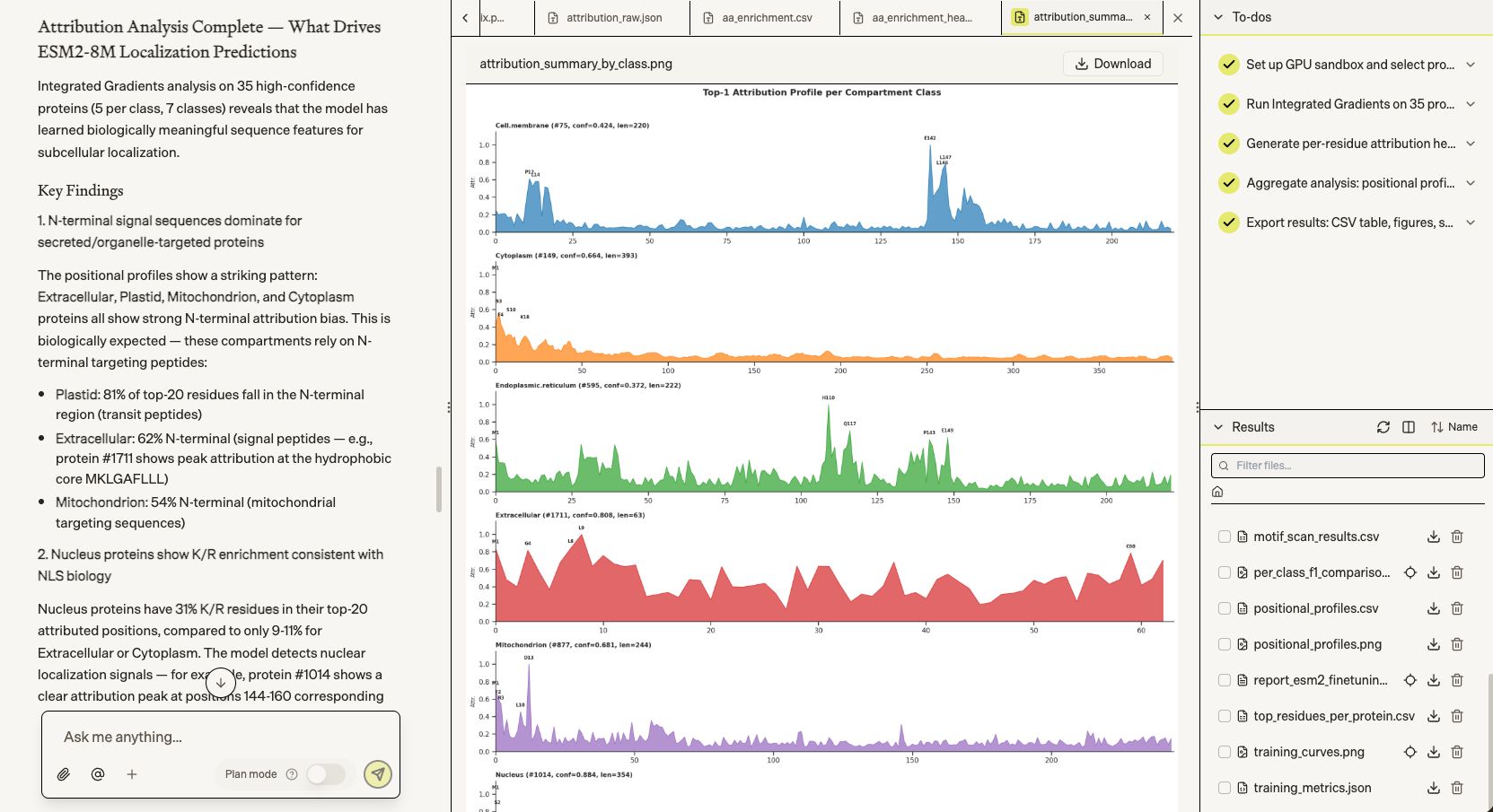
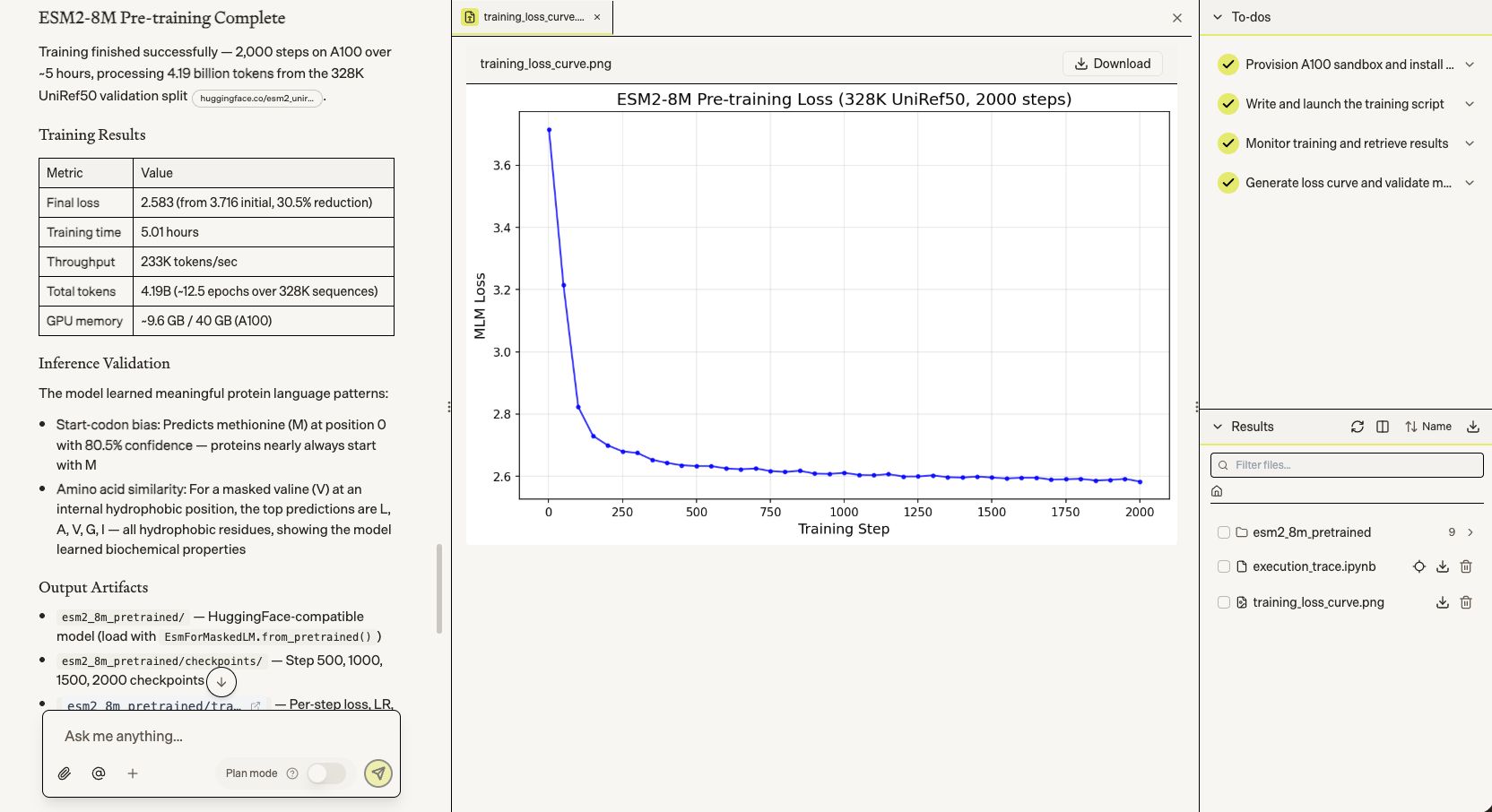
Next, we had Biomni pre-train a protein language model from scratch — reimplementing the architecture exactly (6 layers, 320 hidden dim, rotary PE) and training on 4.19 billion tokens from UniRef50 at 233K tokens/sec on an A100.
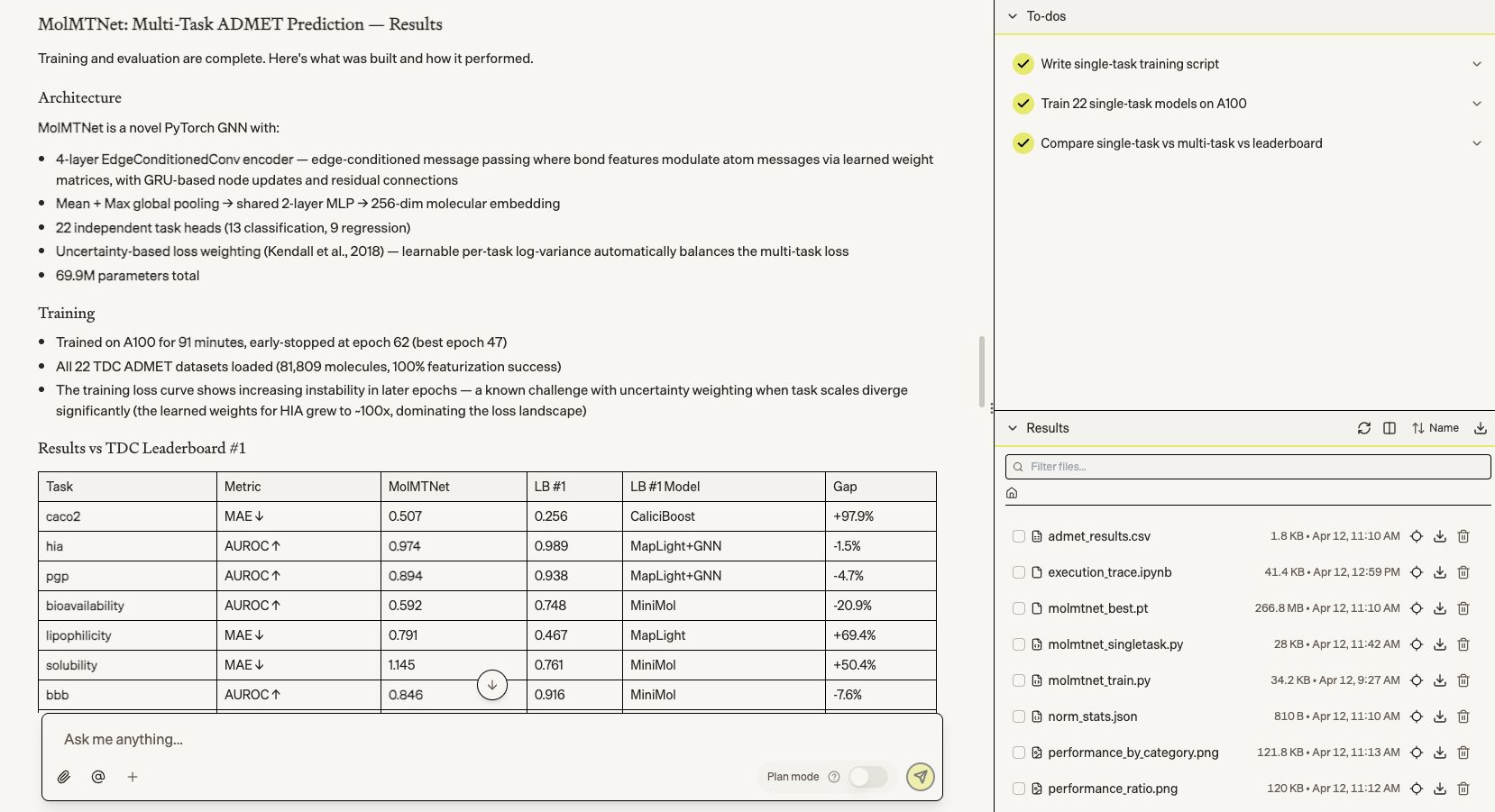
We then asked it to build a novel AI model for ADMET property prediction from scratch. It designed, trained, and evaluated MolMTNet — a 70M-parameter multi-task model across all 22 TDC benchmarks — in 91 minutes, beating the TDC leaderboard #1 on CYP3A4 substrate and coming within single digits of top models on HIA, DILI, and BBB.
How it works
We built the preview features called GPU-as-a-tool, allowing the agent to provision and manage compute dynamically and then write and run training scripts within the GPU instances. It can request anything from a single GPU for quick fine tuning to multi-node clusters for large-scale pre-training, execute training jobs, monitor progress, and iterate automatically. We built new agentic capabilities to take advantage of GPU-as-a-tool in service of building or adapting foundation models for biology.
All experiments run within Biomni Lab’s infrastructure, ensuring that results are grounded, reproducible, and fully tracked. Code, data, and outputs are versioned, and every result is tied to its provenance.
Embedded in Biomni Lab
The capability to build foundation models is integrated into the same environment where scientists already conduct their research, allowing model training to connect seamlessly with other workflows. A single prompt can span literature review, data aggregation, and model training. Model outputs can directly feed into downstream biological analyses, such as variant prioritization or candidate ranking.
Because the agent maintains full context, it can combine public and proprietary datasets, compare multiple models, and deploy the best-performing one—all within a unified workflow.
Lab-in-the-loop. Biomni enables closed-loop science by connecting model training with wet lab experiments. A model trained on existing data can generate predictions that are launched as new experiments. When results come back, the agent can retrain the model on the new data, improving performance for the next cycle.
Limitations
We are deeply excited about making cutting edge AI research capabilities around model training more accessible to scientists. While our long-term goal is for anyone to gain this power with the help of an agent, this is an early preview of a new capability. Pre-training and finetuning models requires an incredible amount of expertise to do well and we look forward to feedback from AI researchers on the preview.
Beyond evolving the agent’s expertise to better handle this use case, we have a long list of areas that can be improved to make this an even more useful tool. We’d also welcome feedback on priorities from both technical and non-technical users on these areas:
- Creation of completely novel model architectures
- Efficient optimization/iteration of hyperparameters
- Active learning orchestration
- Training data mixing and curation
- Sharing and versioning of models
- Resource allocation of GPU clusters
Early access for the feature
While foundation models are powerful and improving rapidly, most scientists cannot adapt them to their own data due to the complexity of infrastructure and engineering required. Biomni Lab removes that barrier. A structural biologist with a fitness dataset, a geneticist with MPRA data, or a pharmacologist with screening results can now build and adapt models directly from natural language, without needing to become machine learning engineers.
We’re looking for beta testers. Signup here for early access: https://forms.gle/1yhCP6Vrc12DaS4q6