Agent-managed sandboxes for scientific workloads
By Zixin Huang, Member of Technical Staff at Phylo
Introducing the manageMachine tool on Biomni Lab, which enables the Biomni agent to dynamically create, delete, and orchestrate sandboxes with configurable compute resources for large-scale biomedical workloads.
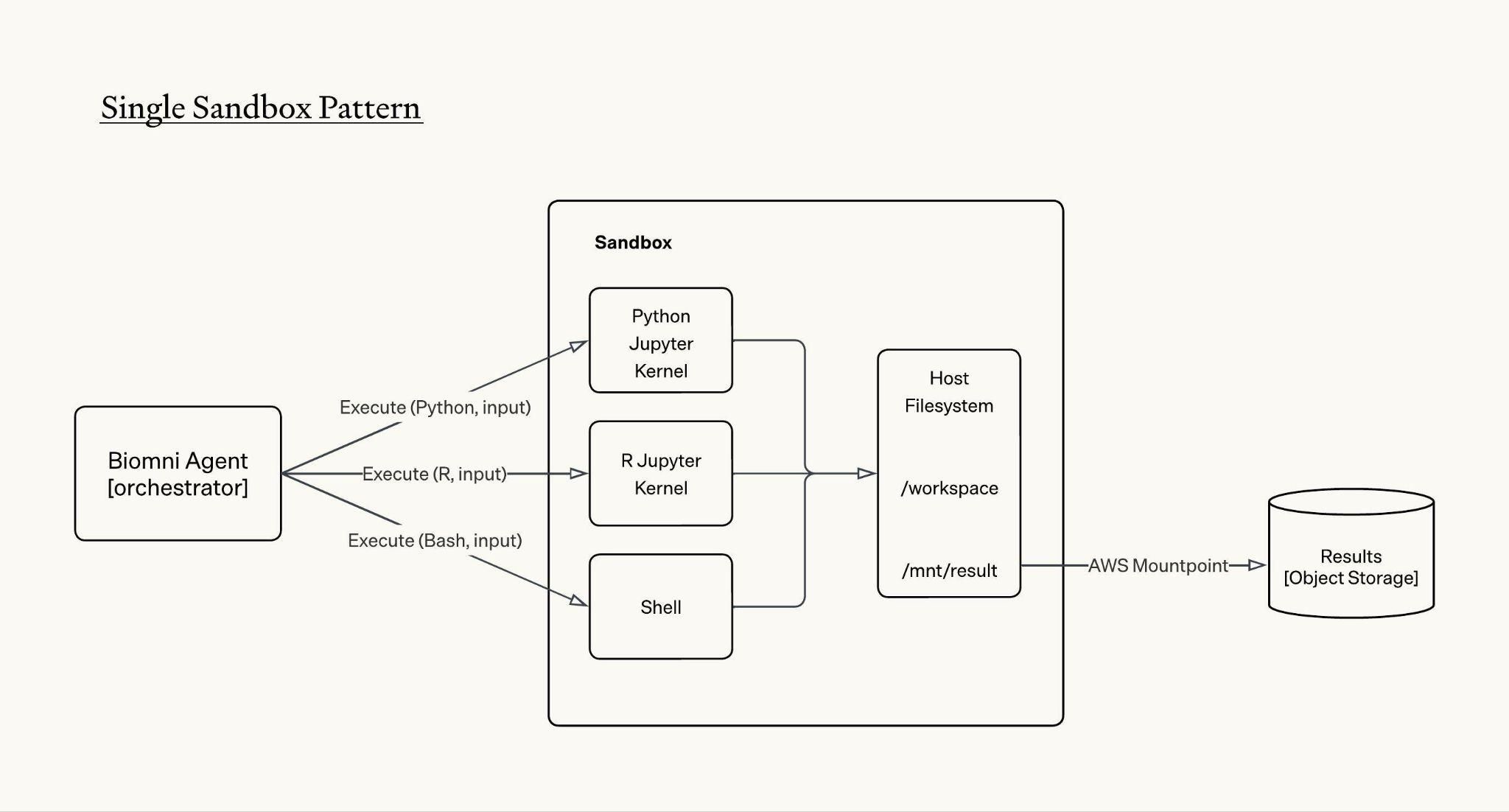
Since Biomni Lab launched three months ago, Biomni agent uses the standard single sandbox pattern to help researchers run diverse biomedical workloads. One piece of feedback we often receive is that when running compute intensive biomedical workloads, the sandbox can be slow, sometimes goes OOM, and gets stuck. We believe the single sandbox pattern is inherently not friendly to biomedical tasks due to the following reasons:
Limited compute resources. Biomedical tasks are memory bound and many of the tasks can be parallelly executed. The single sandbox approach cannot be horizontally scaled and therefore not parallelizable; vertical scaling is also limited as the sandbox still competes for resources on a shared node with a fixed CPU and memory limit, leaving the agent with an often insufficient compute ceiling.
No control over the sandbox’s lifecycle. The sandbox is handed to the agent as a single fixed and programmatically managed environment, meaning when a workload exceeds available resources, the agent has no options to provision, resize, or release compute resources on its own.
As we’ve reflected on how human biologists routinely coordinate with bioinformaticians and infrastructure teams to provision the right compute resources for the task, we realized the Biomni agent should also have the capability to provision compute resources as needed and autonomously manage the lifecycle of compute.
Introducing the manageMachine tool on Biomni Lab which enables Biomni agent to:
- Create new sandboxes with configurable CPU and memory limit, delete running sandboxes, and list metadata of running sandboxes
- Dispatch tool calls and orchestrate tasks across multiple sandboxes
How it works
Behind the scenes, we define the manageMachine tool with the following schema:
tool: manageMachine
params:
action: Create | Delete | List
machines: Array[Machine]
Machine:
name: <machine_id>
type: CPU | GPU
spec: Optional
cpu_cores: int
memory_gb: int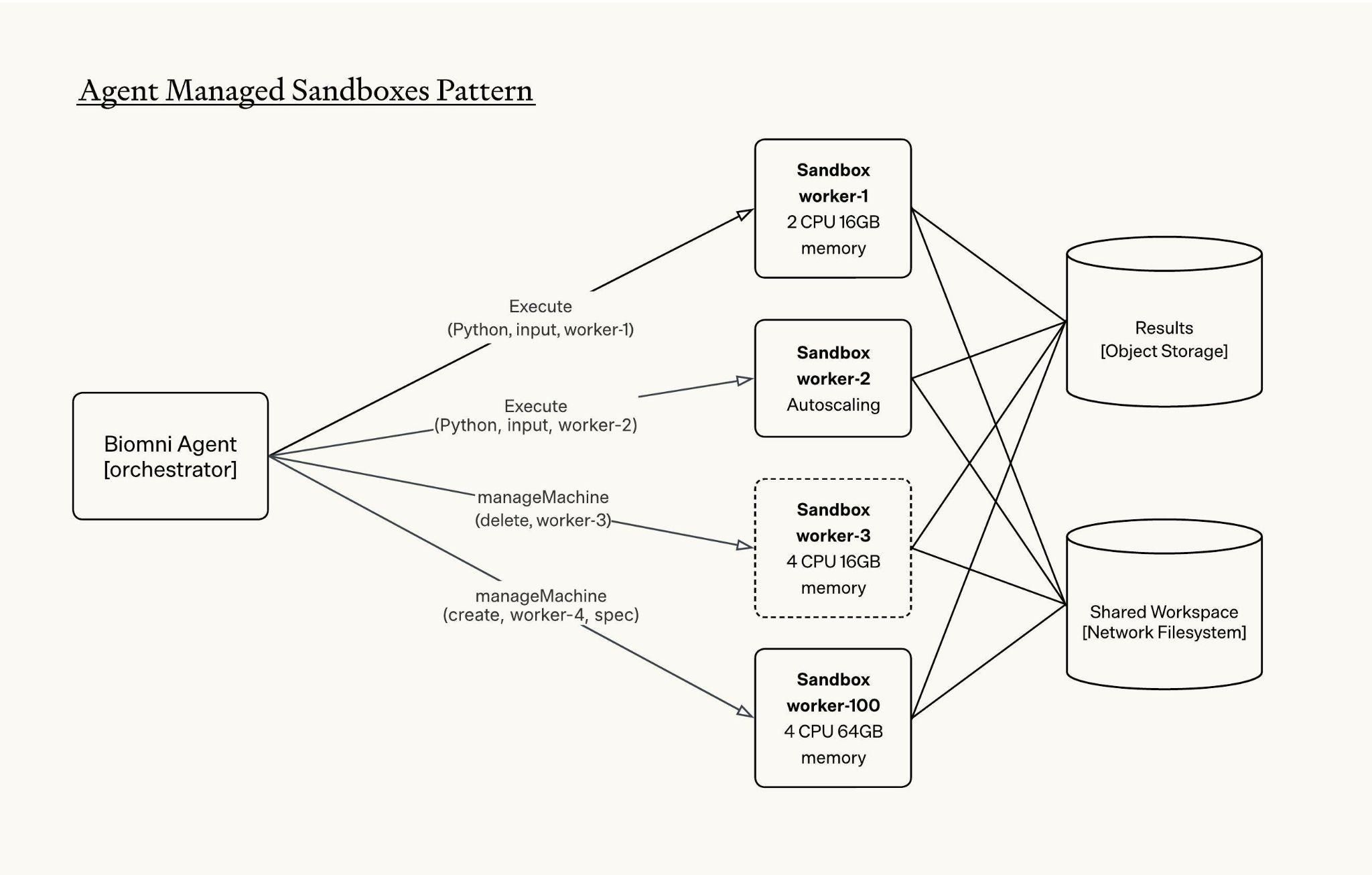
The Biomni agent uses the manageMachine tool to create sandboxes and manage the lifecycles of active sandboxes. The tool also supports bulk sandbox creation, which is critical for biomedical workloads where parallel execution is the norm, and it allows Biomni to spin up multiple instances in a single tool call rather than sequentially.
All sandboxes are mounted to the same network file system, giving them access to a shared workspace. This allows one sandbox to read or use files that were generated by another sandbox.
The agent orchestrates tasks across sandboxes using the unique identifier `machine_id` of each active sandbox. To run bash, code, read, edit and other tool calls, agent specifies which machine-id to send the tool calls to:
execute(tool_name, input, machine_id)An example workflow with agent managed sandboxes would look like:
# create 2 sandboxes with different resource spec
# if spec is not provided, machine is autoscaled and competes for node's resources
1. manageMachine(
action='create',
machines=[
{
name: "worker-1",
},
{
name: "worker-2",
spec: {cpu_cores: 2, memory_gb: 16}
}
]
)
# write some files in worker-1
2. execute(write, string, "worker-1")
# run Python in worker-1
3. execute(Python, code, "worker-1")
# run R code in worker-2
4. execute(R, code, "worker-2")
# run subagent with eligible machines
5. execute(subagent, query, ["worker-1", "worker-2"])
# clean up sandboxes
6. manageMachine(
action='delete',
machines=["worker-1", "worker-2"]
)From one sandbox to many: a new execution model for scientific agents
Giving Biomni the ability to dynamically manage compute resources changes what a scientific agent can do. With a single sandbox, the agent works like a researcher with one laptop: Run one analysis, wait for it to finish, inspect the result, then decide what to do next. That works for small tasks, but biomedical research is often large, parallel, messy, and exploratory by nature. With manageMachine, Biomni instead functions more like a research team. It can create multiple sandboxes, assign each one a different part of the workload, monitor progress, recover from failures, and combine results at the end.
This shift to agent-managed sandboxes creates three immediate benefits.
1. Distributed execution for large-scale biomedical workloads
Biomedical workloads are often naturally distributable. A single RNA-seq analysis can involve multiple stages, such as QC, alignment, quantification, and differential expression, applied across dozens or hundreds of samples where a moderate cohort can involve tens to hundreds of gigabytes of raw sequencing data, intermediate files, and temporary outputs.
With agent-managed sandboxes, Biomni can split sample batches or pipeline stages across multiple sandboxes instead of pushing the entire workload through one constrained environment. This enables faster iteration while also reducing bottlenecks from single-sandbox memory, disk, and runtime limits. For workloads with enough independent branches, hours of sequential execution can become minutes of coordinated execution.
2. Structured exploration
Biomedical analysis often involves choices: which normalization method to use, which statistical model to trust, which gene prioritization strategy to try, or which docking parameters to test. In a single sandbox, the agent usually has to choose one path and commit to it.
With multiple sandboxes, Biomni can fork the workflow: run competing approaches side by side, compare outputs, and converge on the most promising result. This turns the agent’s reasoning into executable experimentation. Instead of only thinking through alternative hypotheses in text, Biomni can test those hypotheses in real compute environments.
3. Agent as fault tolerance mechanism
Real bioinformatics pipelines fail often. Dependencies conflict, file formats are inconsistent, tools crash, and some jobs silently produce bad outputs.
The agent can remain in the loop as a fault-tolerance layer: monitoring sandbox failures, detecting incomplete or corrupted outputs, retrying failed samples, adjusting resource allocation, and resuming only the affected branches rather than restarting the entire analysis.
Together, these capabilities make Biomni more than a single-environment executor. At a system level, this is the foundation for agent-managed infrastructure. The agent is no longer limited to the one machine it was given. It can provision resources, isolate work, distribute execution, manage failures, and optimize across speed, cost, and accuracy.
Looking ahead
Scaling to hundreds of sandboxes
Internally we’ve been experimenting with agents orchestrating hundreds of sandboxes within a single session. Scaling to that level introduces a new bottleneck: Coordination between sandboxes becomes the hard part, not the computation itself.
One architectural improvement we are exploring is a coordinator-worker pattern. The agent spins up a dedicated coordinator sandbox responsible for maintaining shared data structures, such as task queues, state registries, and result buffers, while worker sandboxes focus purely on computation. This separation of orchestration from execution gives the agent a reliable coordination layer, reducing errors and improving reproducibility at scale.
Environment as a primitive
Another direction we are exploring is making execution environments a first-class primitive in manageMachine.
Biomedical tools often have conflicting requirements: specific package versions, system libraries, CUDA versions, compiled dependencies, or container images. A single sandbox can run many tools, including both R and Python, but not every workload belongs in the same environment.
To support this, we can extend the Machine schema with an image field:
Machine:
name: <machine_id>
type: CPU | GPU
image: Optional[str]
spec: Optional
cpu_cores: int
memory_gb: intFor more dynamic workflows, we can also imagine a createCustomImage tool that lets Biomni build a new environment from a base image and a set of dependencies:
tool: createCustomImage
params:
name: <image_name>
dockerfile: <dockerfile_content>Conclusion
Agent-managed infrastructure is still in its early stages, but the direction is clear. Over time, we envision Biomni managing even more of the execution layer: larger sandbox pools, coordinator-worker workflows, custom environments, and eventually full compute clusters. Our goal is to empower Biomni with the same tools and infrastructure that human researchers can access today. By making compute infrastructure native to the agentic scientific workflow, we can accelerate autonomous research for every biomedical scientist.