back

Cross-Species Validation of Mouse Model Findings Using Human Single-Cell Data
"I consider myself a veteran in single-cell genomics, yet having an expert-level companion like Biomni Lab has been genuinely transformative. I used to verify every step manually, but Biomni Lab consistently delivers reliable, high-confidence results while dramatically reducing the time required for dataset discovery and analysis. This allows me to focus on generating biological insight rather than managing technical workflows." — Xiaochen Xiong, Biology PhD student at Stanford University
Mar 25, 2026
Pancreatic ductal adenocarcinoma (PDAC) is among the most lethal solid tumors, and a major reason is that we still understand relatively little about its microenvironment. Recent work has shown that the nervous system plays an active role in shaping tumor progression, not simply as a bystander but as a participant in the signaling that drives disease. These neuro-tumor interactions represent a promising but underexplored area for therapeutic intervention.
Xiaochen Xiong, a PhD student in the Monte Winslow lab and Marc Tessier-Lavigne lab at Stanford, has been studying the molecular crosstalk between pancreatic tumors and sympathetic neurons. Through a series of mouse experiments, he pinpointed the NPY (Neuropeptide Y) signaling pathway as a key mediator of this interaction. The natural next step was to ask whether the same biology holds in human tissue.
The translational gap between mouse and human
Validating a mouse finding in humans is conceptually straightforward but technically demanding. For the NPY pathway, this meant characterizing an entire signaling axis across human cell types: NPY itself, its receptors (NPY1R, NPY2R), the processing enzymes that activate it (PCSK1, PCSK2), and the downstream G-protein signaling molecules. Each of these could have a distinct expression pattern across sympathetic neurons, epithelial cells, fibroblasts, endothelial cells, and immune cells in the human lung and nervous system.
In practice, this kind of cross-species validation involves a long chain of bioinformatics work: finding the right publicly available human single-cell RNA-seq datasets, dealing with inconsistent storage formats across repositories, writing scripts to download and process the data, running quality control and normalization, and generating visualizations that are actually interpretable. The process commonly takes weeks or months before anyone can even begin looking at biology.
Cutting through the work with Biomni Lab
Rather than going through that pipeline manually, Xiaochen used Biomni Lab. Through a short series of natural-language prompts, and guided by Biomni Lab's own follow-up questions to refine the analysis, he worked through the key questions driving the project.
“Are there single-cell sequencing data for human sympathetic neurons? ”
Biomni Lab searched the literature and public repositories, identified high-quality single-cell datasets for sympathetic ganglia and retrieved the data autonomously.
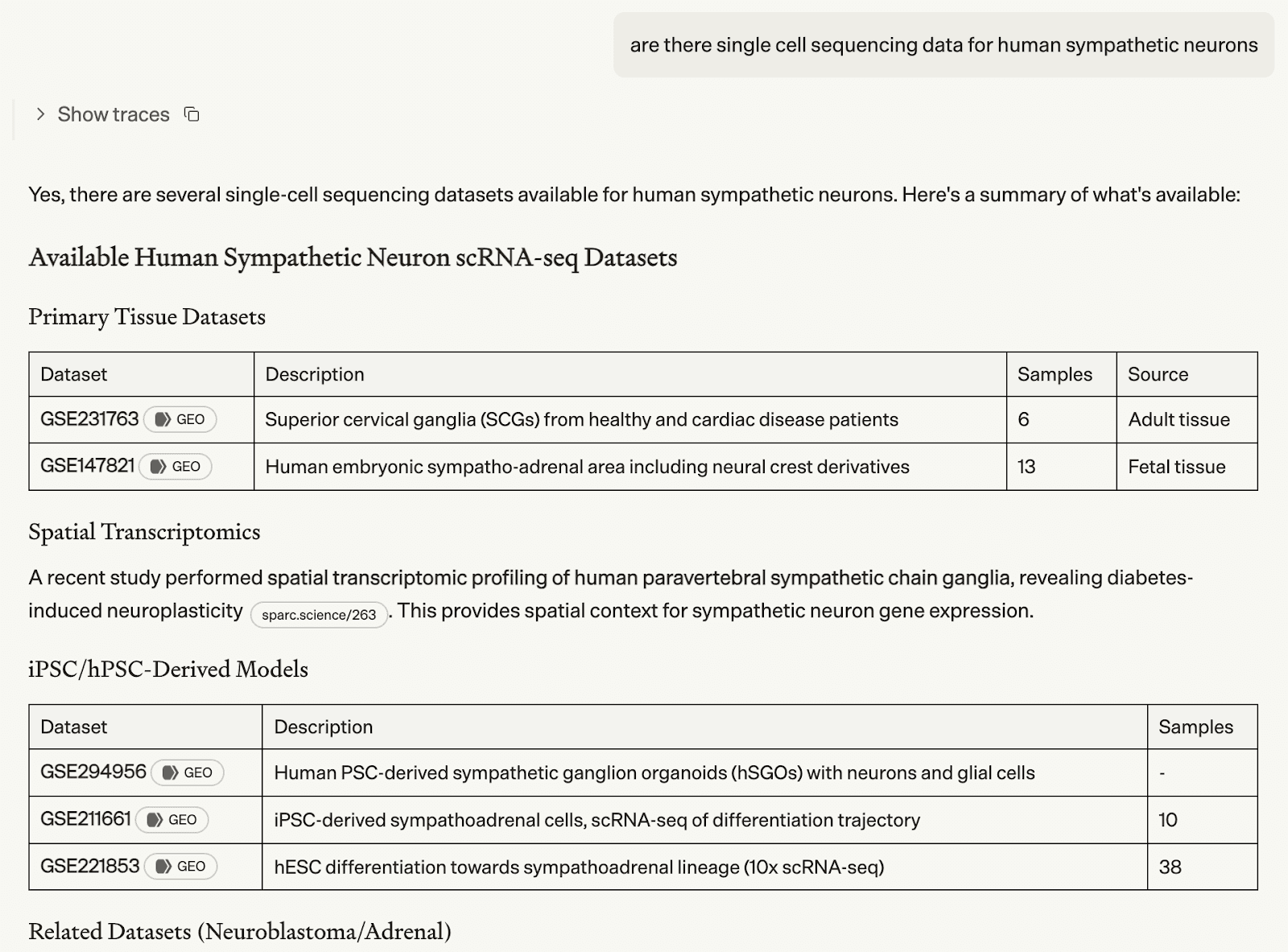
Figure 1. Screenshot of Biomni Lab surveying available human sympathetic neuron scRNA-seq datasets across public repositories.
“How are NPY pathway components distributed across human sympathetic neuron subtypes?”
Biomni Lab generated UMAP plots delineating the cellular landscape of human sympathetic ganglia and mapped the expression of NPY, its receptors, processing enzymes, and signaling molecules across each subpopulation. The resulting expression profiles showed which neuron subtypes are the primary sources of NPY signaling, providing a direct comparison to the mouse data.
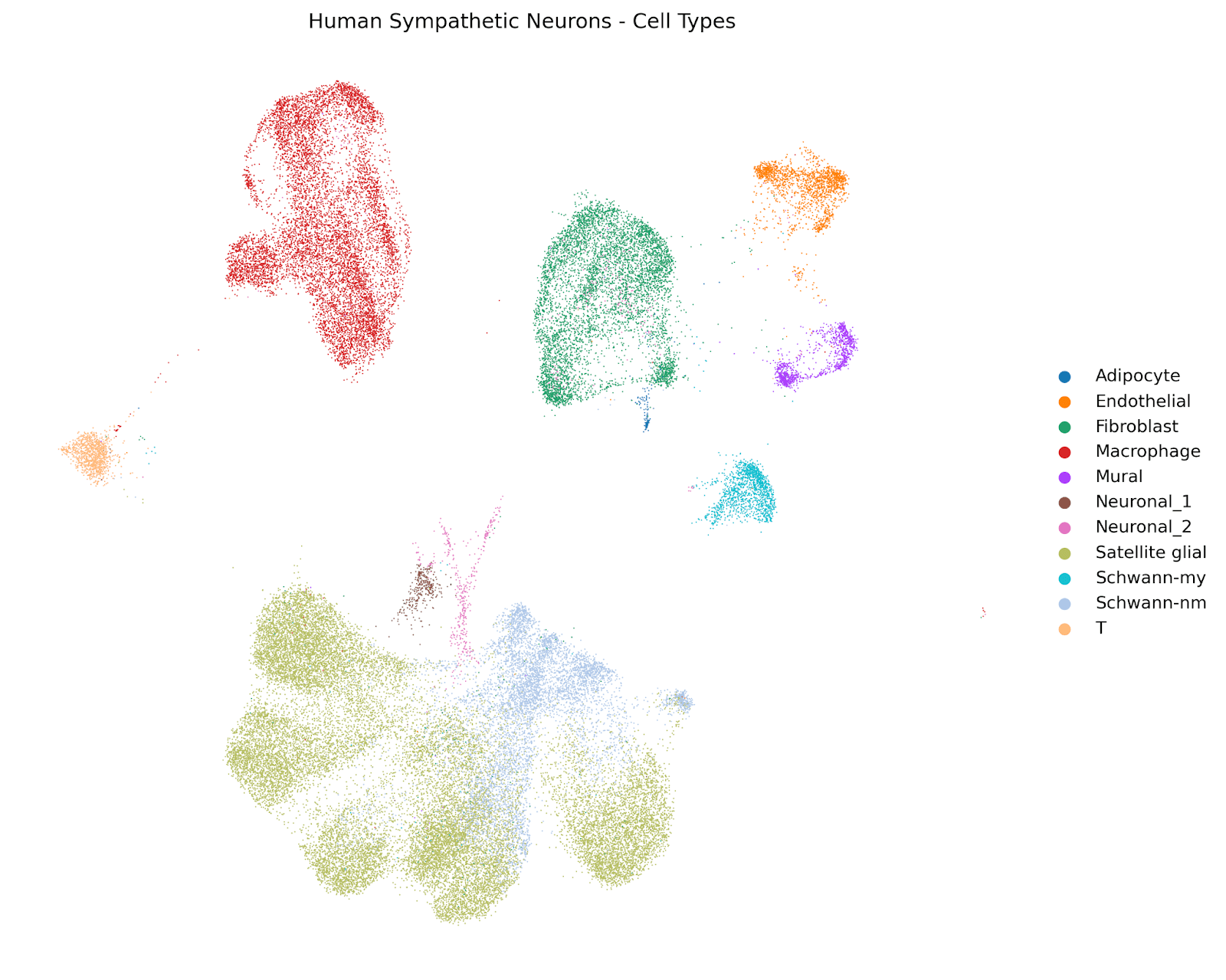
Figure 2. UMAP visualization of human sympathetic ganglion cell types from single-cell RNA sequencing.
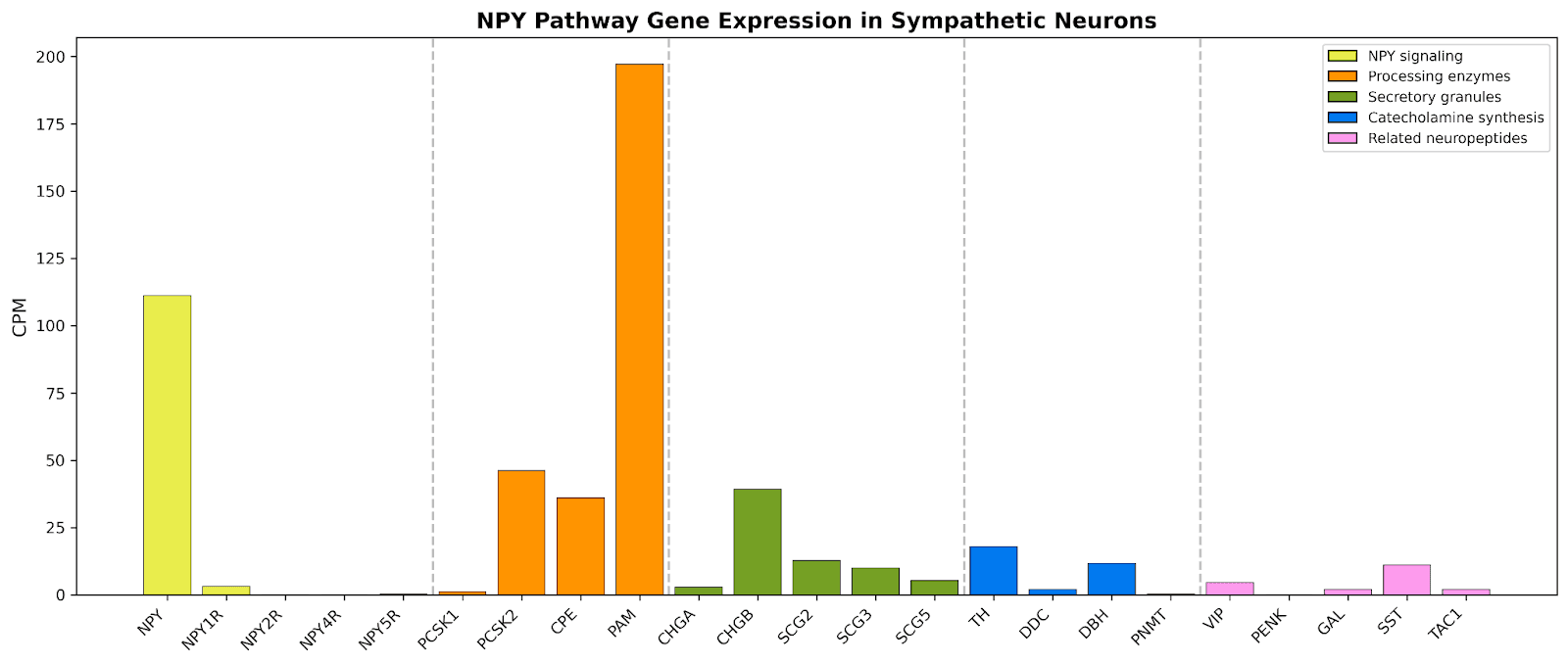
Figure 3: Expression profile of NPY pathway genes in human sympathetic neurons.
“Does the lung microenvironment express the complementary receptors and ligands needed for the signaling axis to function?”
Biomni Lab profiled NPY pathway gene expression across lung cell types, including sympathetic neurons, blood cells, epithelial cells, endothelial cells, and fibroblasts. The output included detailed heatmaps and barplots showing where the pathway's components converge.
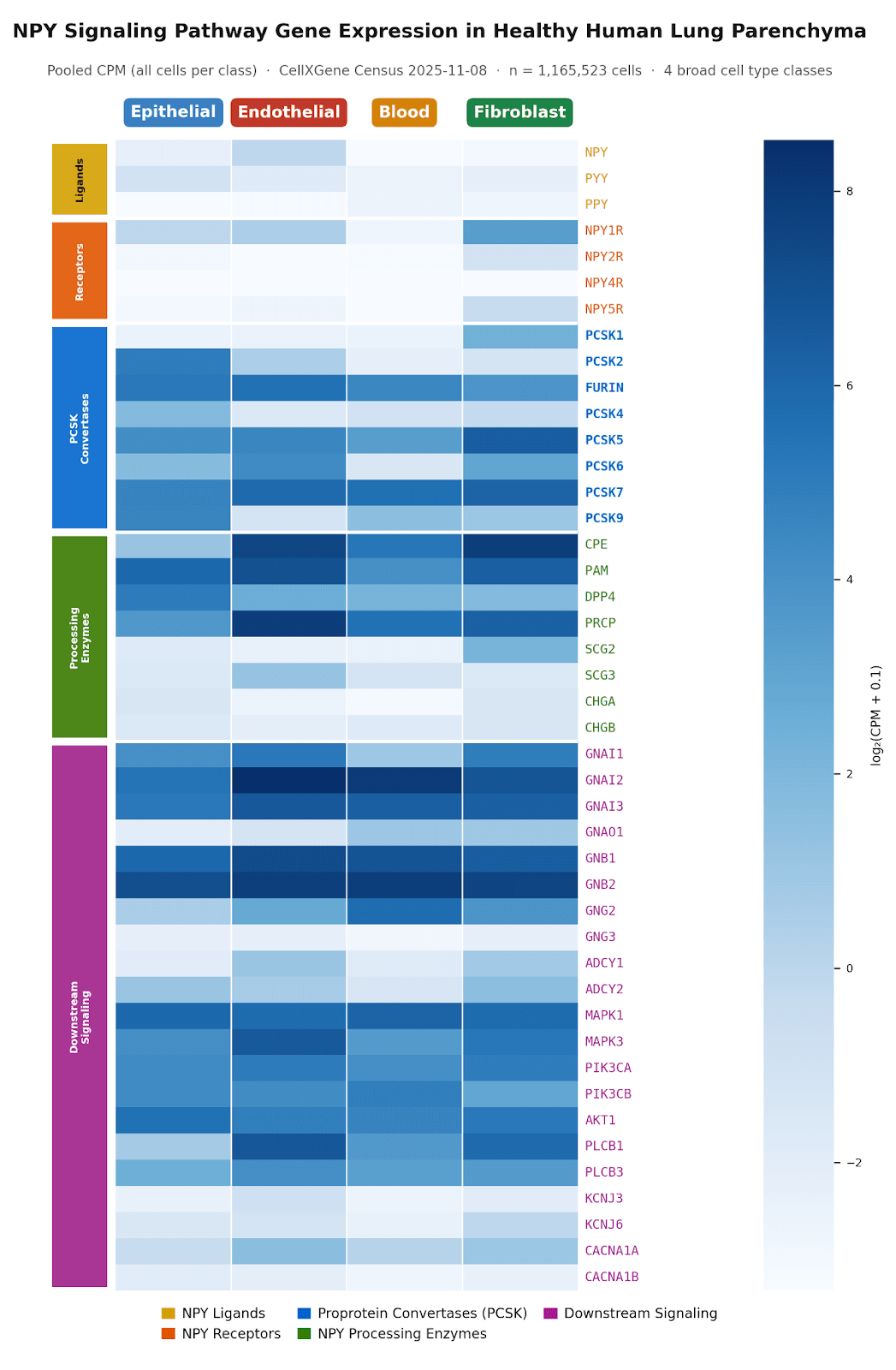
Figure 4: NPY pathway gene expression across human lung cell types (blood, epithelial, endothelial, fibroblast).
The full analysis, from dataset discovery through presentation-ready figures and a standardized data table for downstream statistics, was completed in a fraction of the time the conventional approach would have required.
Bringing the mouse and human pictures together
The human single-cell data confirmed what the mouse model had suggested. NPY pathway components showed clear, cell-type-specific expression patterns in both human sympathetic ganglia and lung tissue, consistent with the signaling architecture the Xiaochen had characterized in mice. This cross-species concordance supports the interpretation that the NPY axis is a conserved biological mechanism with real clinical relevance, one that extends beyond the mouse model. It also strengthens the rationale for exploring this pathway as a potential therapeutic target.
Looking ahead
Much of the time in computational biology is spent on logistics: finding the data, reformatting it, running standard processing steps. These are necessary but they aren't where scientific thinking happens. By taking over that work, Biomni Lab lets researchers stay focused on the biology itself, on interpreting results, forming hypotheses, and designing the next experiment.
For the Winslow and Tessier-Lavigne labs, this means faster cycles between mouse discoveries and human validation, and more time for the work that actually moves the science forward.