back

Scaling Agentic Clinical Trial Emulation Using Real-World EHR Data at Mount Sinai
Mount Sinai researchers used Biomni to autonomously scale target trial emulation across multiple landmark anticoagulation studies, turning months of manual EHR analysis into repeatable, agent-driven workflows. By enabling high-volume, standardized trial replication, Biomni made it possible to systematically learn how published RCT results translate within a local care environment—shifting trial emulation from one-off analyses to scalable institutional insight.
Feb 22, 2026
Mount Sinai researchers used Biomni to autonomously scale target trial emulation across multiple landmark anticoagulation studies, turning months of manual EHR analysis into repeatable, agent-driven workflows. By enabling high-volume, standardized trial replication, Biomni made it possible to systematically learn how published RCT results translate within a local care environment—shifting trial emulation from one-off analyses to scalable institutional insight.
We're excited to share how researchers at Mount Sinai's Icahn School of Medicine are using Biomni to autonomously emulate clinical trials against real-world patient data, turning months of manual analysis into repeatable, agent-driven workflows.
Trial Evidence Doesn't Always Translate Locally
Randomized clinical trials (RCTs) remain the gold standard for measuring drug efficacy, but when a hospital system asks whether a drug works the same way for their patients, the answer is rarely straightforward. Differences in patient demographics, prescribing patterns, adherence, and how outcomes get recorded can all cause real-world evidence to diverge from published trial results.
This gap matters. When clinicians choose between treatments, say a newer anticoagulant versus warfarin for atrial fibrillation patients, the relevant question goes beyond what a landmark trial found. What they really need to know is how that effect plays out in their institution, with their patient population.
The standard approach to bridging this gap is target trial emulation, which replicates an RCT's design using electronic health record (EHR) data. But these emulations are labor-intensive. They require protocol interpretation, phenotype construction, cohort assembly, confounder adjustment, survival modeling, and extensive diagnostics. Most institutions can manage only a handful, which is far too few to learn systematic patterns about how trial evidence behaves locally.
Reading the Signal in Trial Discrepancies
A team of researchers at Mount Sinai, led by Justin Kauffman, Joshua Lampert, MD and Benjamin Glicksberg, PhD proposed a fundamentally different framing: What if the discrepancies between trial results and real-world evidence aren't noise to be minimized, but structured information worth learning from?
Their hypothesis was that an institution's pattern of agreement and disagreement with published trials encodes something real, a "transport relationship" that reflects how the local care environment systematically transforms clinical evidence. Testing that hypothesis requires scale. You can't learn a systematic pattern from one emulation. You need many, executed in a standardized and reproducible way.
That's where Biomni comes in.
“As someone who’s worked in EHR-based analyses for years, I’ve learned that generalizability is earned across the full pipeline and that small sources of fragility can accumulate at every step: cohorting, concept sets, phenotypes, confounding control, diagnostics, and iteration. Agents like Biomni can finally bridge those gaps and scale trial emulation in ways we couldn’t realistically do by hand. But as with any new innovation, we need to pressure-test these systems with rigorous, transparent evaluation before deploying them in practice. Biomni has already made waves in computational biology, and this is the first step toward doing the same for real-world evidence in EHRs and beyond.”
- Dr. Benjamin Glicksberg
How Biomni Powers End-to-End Trial Emulation
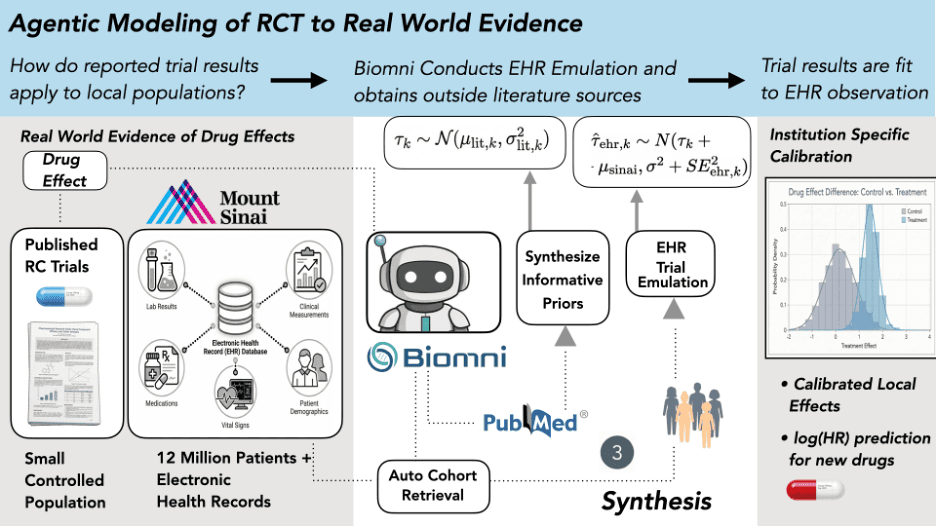
Biomni served as the autonomous engine for the entire emulation pipeline. Given a set of written instruction documents and access to Mount Sinai's OMOP-mapped EHR database (covering 12 million patients), Biomni independently executed each stage of the workflow:
Protocol Parsing — The agent interpreted published trial protocols and extracted eligibility criteria, treatment definitions, and endpoint specifications.
OMOP Concept Set Construction — Biomni mapped clinical concepts (drugs, diagnoses, procedures) to standardized OMOP vocabulary codes, a task that typically requires deep domain expertise and hours of manual curation.
Cohort Building & Covariate Extraction — The agent assembled treatment and control cohorts from the EHR, identified relevant confounders, and extracted baseline covariates for adjustment.
Confounder Adjustment & Effect Estimation — Biomni applied propensity score methods, fit survival models, and estimated treatment effects on the log-hazard ratio scale.
Literature Synthesis — Beyond the EHR, Biomni conducted structured literature searches to build comparison-specific priors, quantifying how much EHR–RCT disagreement is typically observed for each drug comparison based on published evidence.
Discrepancy Diagnosis & Refinement — When initial results flagged quality issues, the agent diagnosed potential problems and, where indicated, refined and re-ran the emulation.
Crucially, this entire pipeline ran three independent times per trial—each starting from the same instructions but making its own analytic decisions, to quantify the variability introduced by the agent's degrees of freedom. This turned a potential weakness of AI-driven analysis (stochastic decision-making) into a measurable source of uncertainty.
What the Emulations Found
The team emulated five atrial fibrillation anticoagulation trials across Mount Sinai's EHR: ARISTOTLE (apixaban), ROCKET AF (rivaroxaban), RE-LY (dabigatran), ENGAGE AF–TIMI 48 (edoxaban), and AVERROES (apixaban vs. aspirin).
Biomni's emulations, combined with a Bayesian hierarchical calibration model, produced striking results:
100% coverage of published trial effects — All four held-out trial results fell within the model's 95% posterior predictive intervals during leave-one-out cross-validation.
A consistent institutional signal — The model identified a stable, positive institutional shift (μ_Sinai), indicating that DOAC benefit was systematically attenuated in Mount Sinai's EHR relative to published trial results, likely reflecting high-quality warfarin management, local prescribing patterns, and outcome ascertainment differences.
Out-of-distribution generalization — When applied to AVERROES (a structurally different trial comparing apixaban to aspirin rather than warfarin), the model reduced prediction error by 86.5%, suggesting it learned a genuine system-level property, not a comparator-specific artifact.
Scaling Single Emulations to Institutional Learning
Traditional trial emulation asks whether we can reproduce a given trial result. The Mount Sinai team, powered by Biomni, pursued a deeper question: what does the pattern of agreement and disagreement across many trials tell us about how their institution transforms clinical evidence?
That shift from isolated emulations to cumulative institutional learning only becomes possible at the scale that autonomous agents enable. A single emulation is essentially a case report. But when you run many standardized emulations and analyze them jointly, you start to characterize how clinical evidence actually behaves within a specific care environment.
For Biomni, this work points toward a new paradigm for AI agents in clinical research. The real contribution is generating the volume of standardized, reproducible analyses needed to surface system-level properties that no single study could reveal.
What's Next
This research represents one example of what becomes possible when clinical research workflows can run at scale. We see growing opportunities for autonomous agents across clinical evidence generation, from trial emulation and outcome benchmarking to safety signal detection and comparative effectiveness research. As more institutions bring these tools into their research environments, the scope of questions they can ask will expand well beyond what manual analysis pipelines allow.
The Mount Sinai team is continuing to push this work forward, with plans to extend their framework to safety endpoints and cross-institution learning.
Preprint is now available here.
Try Biomni Lab today.