Evaluating AI Agents in Biology
As AI agents become central to biological research, evaluation must keep pace. We examine why existing benchmarks fall short for biology, share lessons from our experience with BixBench including a verified subset, and introduce BiomniBench, a trace-based evaluation framework that scores agents on their analytical process, not just the final answer. Biomni Lab achieves state-of-the-art performance across both general-purpose and domain-specific agents on both benchmarks.
Evaluation Matters for Biology
Imagine an AI agent analyzing a differential expression dataset. It returns a clean volcano plot, a ranked gene list, and a confident interpretation. Everything looks right. But buried in the trace, it used the wrong normalization, ignored batch effects, and cited a paper that doesn't exist.
This is the core problem with biology agents: failures are silent. In software, bad code crashes. In math, an invalid proof is verifiable. In biology, a flawed analysis keeps moving. It informs wet lab experiments, shapes drug development decisions, and can propagate through a research pipeline for months before anyone notices. The cost is material—it's wasted reagents, failed trials, and conclusions built on a foundation that was never sound.
So how do we know these agents are actually working? Evaluation. But building good evals for biology is a genuinely hard problem.
Why Evaluation for Biology Is Uniquely Difficult
Biology is different from other domains for several reasons.
Diverse domains, diverse tasks. Biology is not one homogeneous field; it spans many complex domains, including molecular biology, genomics, ecology, structural biology, clinical research, and dozens of other sub-domains, each with its own methods, terminology, and standards. On top of that, the task modalities are wildly varied. A biologist (or an agent) may need to review and synthesize literature, interpret microscopy images, parse genomic data, read figures, and write protocols. Building evaluation coverage across this space is a challenge.
Biology doesn't happen in one step. A biologist rarely does one thing in isolation. A typical workflow chains together literature search, experimental design, data processing, analysis and interpretation. Most existing evaluations test these capabilities individually, but real biological work requires getting the entire sequence right.
There's often no single right answer. The messiest fact of all is that there are often multiple valid experimental designs, analytical choices, and interpretations of the same data. Evaluation frameworks that expect one ground truth answer struggle to capture the nuanced reasoning of scientists.
Verification is slow and expensive. Some tasks can only be truly validated by running an experiment, which takes days to months. Even evaluating whether each step was done correctly requires deep domain expertise. A plausible-sounding analysis can be subtly wrong in ways only a domain expert would catch, for example, the wrong statistical test for a data distribution, an inappropriate reference genome, or a confounded comparison. Even early career scientists make these errors.
Existing Benchmarks for Biology
The community has made meaningful progress. Over the past two years, a growing number of benchmarks have emerged to measure AI capabilities in biology.
Early efforts like HLE and LAB-Bench focused on biological knowledge and specific LLM capabilities, such as literature comprehension, figure interpretation, sequence manipulation, and protocol reasoning. More recently, agent-oriented benchmarks have pushed evaluation toward open-ended, multi-step work. Biomni-Eval1, BixBench and others all ask agents to perform real analytical tasks, build pipelines, or make decisions with actual biological data.
These benchmarks reflect real effort and have sharpened our understanding of what LLMs and agents can do in biology. If we want to assess how much LLMs and agents can truly facilitate discovery, the next step is to begin closing the gap between the tasks the existing benchmarks evaluate and the daily workflows of practicing biologists.
Learnings from an Existing Benchmark
Last week, we announced Biomni Lab, our new Integrated Biology Environment. To evaluate its analytical capabilities, we first worked with BixBench, a benchmark where agents analyze real biological datasets and answer short-answer research questions spanning a range of bioinformatics domains. We believe it is one of the most realistic benchmarks available today for assessing data analysis and interpretation, and it has been adopted by other groups, making it a natural starting point.
Our Results
We evaluated Biomni Lab alongside other domain-specific and general-purpose agents on the full BixBench benchmark. Biomni Lab (2026-02-03) led at 52.2% accuracy, followed by Edison Analysis at 42.4%, Claude Code (Opus 4.6) at 39.5%, and OpenAI Agents SDK (GPT-5.2) at 38.5%.
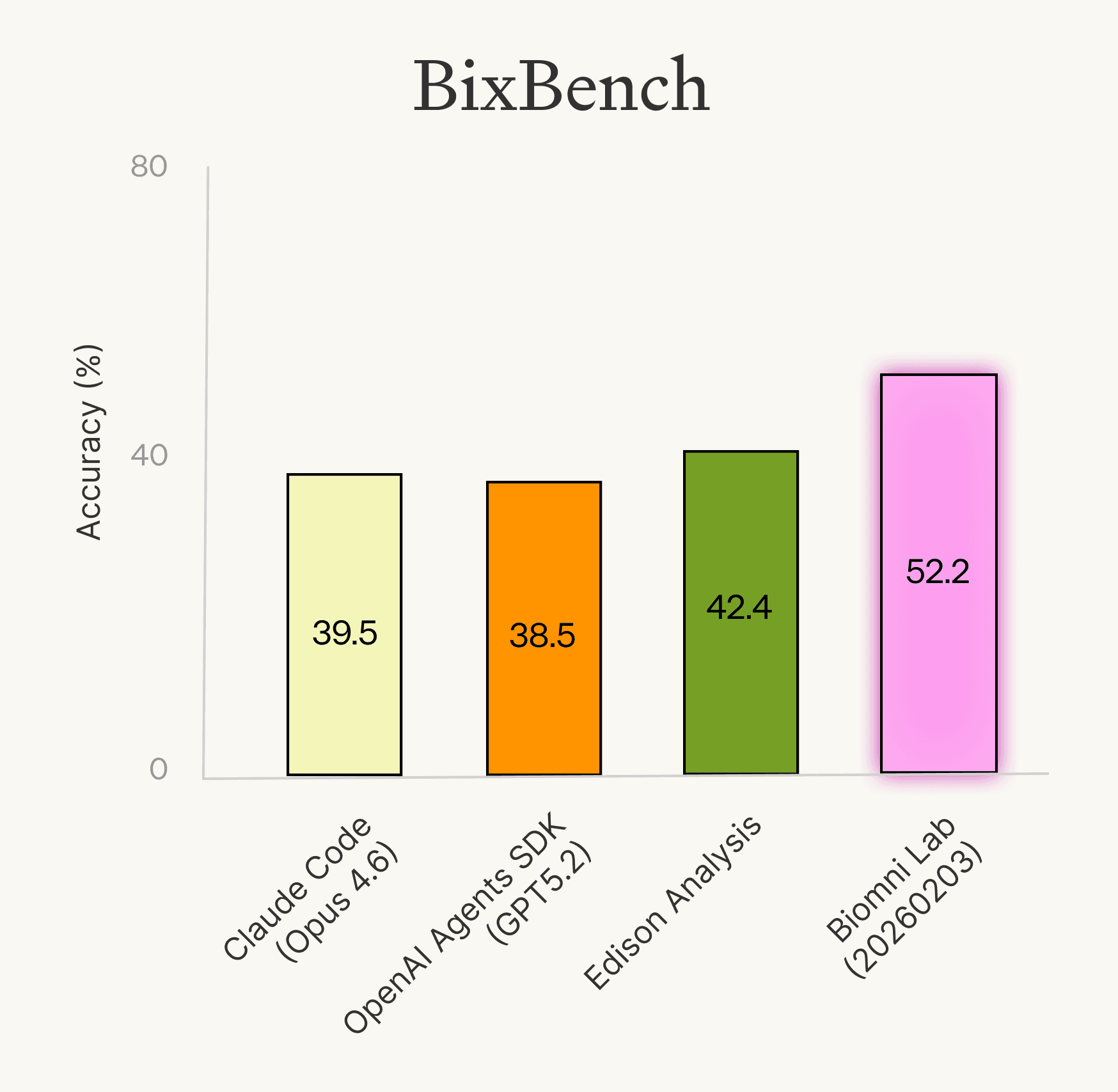
These scores represent meaningful progress from the original paper's ~21% accuracy, but taken at face value, they paint a discouraging picture: even the best agent gets roughly half the questions wrong, suggesting these systems are still far from being practically useful. That doesn't match what we experienced and heard from users, who found them genuinely helpful in their day-to-day research. We wanted to understand this disconnect better, so we looked more closely at what was actually going wrong.
Not All Failures Are Agent Failures
When we dug into why agents got questions wrong, we found the failures fell into distinct categories:
Lack of biological knowledge. The agent lacked a deep understanding of biological concepts needed to perform the analysis correctly. For example, agents struggled with the directionality of biological concepts, mixing together upregulated and downregulated genes during pathway analysis, or misinterpreting the direction of gene essentiality scores. These are real gaps in understanding that are expressed in the agent’s capabilities.
Ambiguous or underspecified questions. The question or context didn't contain enough information for a competent agent, or even a human in many cases, to arrive at the single expected answer. For instance, some questions didn't specify whether "benign" should include both Benign and Likely Benign sample categories, or whether pathway enrichment should use GSEA versus ORA, or what output format was expected.
Incorrect ground truth. The benchmark's reference answer was itself flawed. In some cases, the ground truth analysis ran DESeq2 on already-normalized data, removed samples based on unstated discretion, or used a different file than the one specified in the question.
Only the first category reflects a genuine limitation of the agent. The other two are benchmark failures, not agent failures. They are a reminder that evaluating the eval is itself an essential step.
BixBench-verified-50
To isolate real agent performance from benchmark issues, we curated BixBench-Verified-50. Starting from the full benchmark, we sampled questions and identified problematic ones. Some were removed entirely. For others, we revised the question text for clarity or corrected the expected answer, while being careful not to overcorrect: we used our best judgment to leave in questions where we believe a competent expert should be able to fill in the details themselves. Questions were reviewed together with several domain experts to ensure correctness of ground truth, sufficiency of provided context, and clarity of the expected answer. The verified subset is available on Hugging Face (link), along with a detailed sheet documenting the revisions made and the underlying issues found in each question.
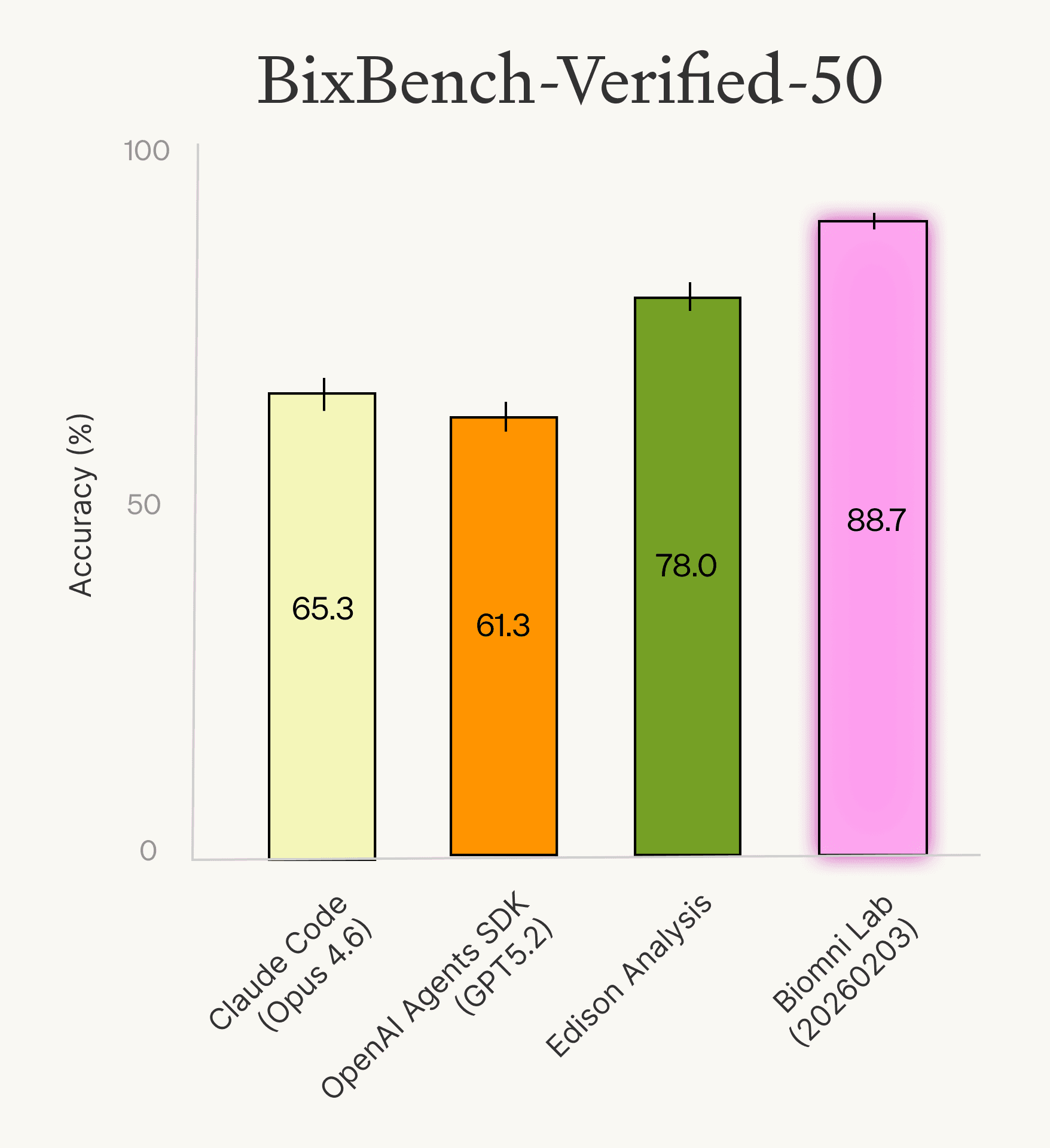
We re-ran the same agents on this verified subset and found the results tell a clearer story on the current state of agent performance. Biomni Lab (2026-02-03) led at 88.7%, followed by Edison Analysis at 78.0%, OpenAI Agents SDK (GPT-5.2) at 61.3%, and Claude Code (Opus 4.6) at 65.3%. Accuracy jumped significantly across all agents, confirming that a meaningful portion of "failures" on the original benchmark were due to benchmark quality rather than agent limitations.
A Deeper Problem
The BixBench exercise exposed a fundamental limitation of using binary scoring for evaluation of agents (a practice common across many of the existing benchmarks). For short factual questions, this works fine. For the kind of long-horizon analytical tasks that define real biological research, a 0-or-1 score throws away most of the (really valuable!) evaluation signal that helps us make research progress.
Consider an agent that loads the right data, applies appropriate filters, then chooses a reasonable but different statistical method, leading to a different result than the ground truth. Under binary scoring, this agent receives the same zero as one that failed at every step. The score reveals nothing about where the agent fell short or what it did well.
Two Ways to Think About Evaluation
The result: "Did you get the right answer?"
This is the most intuitive form of evaluation. Given a task, does the agent produce a correct final output? A correct gene list. A valid protein structure. A solid experimental protocol. Most existing benchmarks operate in this mode: compare the agent's output to a ground truth and score accordingly.
Result-based evaluation is valuable. It scales well, produces clear metrics, and tells us something real about capability. But it gives you very little insight into what the agent actually did. It cannot distinguish between sound reasoning and a flawed process that happened to produce the right answer. It struggles with questions where multiple conclusions or interpretations are equally valid. Without visibility into the process, it becomes difficult to diagnose failures, guide improvements, or build trust in the system.
The trace: "Did you do it in the right way?"
This is how biologists often evaluate rigor. Did you use an appropriate statistical test? Did you account for confounders? Did you cite reliable sources? Did your reasoning hold up at each step?
Trace-based evaluation examines the process, not just the final results. It asks whether the agent chose the right tools for the task, relied on credible and up-to-date references, followed a sound chain of analytical decisions, and applied statistical and computational methods that were appropriate for the data.
This mirrors how science itself works. Peer review doesn't stop at whether a paper's conclusions seem right, in part because experimental verification is often slow, expensive, and impractical at review time. Instead, reviewers scrutinize the methods, the analytical choices, the controls. A PI doesn't just look at a trainee's final figure. They ask "why did you choose this normalization?" and "what happens if you remove that outlier?" Process is where trust is built, precisely because confirming results directly is so costly.
Trace-based evaluation for biology agents is largely missing today. We think that's a critical gap.
BiomniBench: First Trace-Based Evaluation for Biology Agents
Ground truth issues aside, we believed it was imperative to build something new to address the significant limitations of binary scoring for biology tasks.
BiomniBench is our effort to create a trace-based evaluation framework for biology agents. We envision it covering, over time, the range of real-world tasks biologists face: data analysis and interpretation, experimental planning, protein design, and others. We are starting with data analysis, which we call BiomniBench-DataAnalysis. Data analysis and interpretation is one of the most common and important tasks in biological research, and one where agents are increasingly being used.
Design Philosophy
Evaluate the process, not just the output. We score agents on the quality of their analytical decisions at each step: whether they loaded the right data, filtered it correctly, chose appropriate methods, and interpreted the results soundly.
Ground evaluation in real-world tasks. Tasks are long-horizon analyses derived from published high-impact papers, covering both academic and industry research.
Cover breadth across biology. Tasks span different biomedical domains and data modalities, reflecting the diversity of analytical work biologists face every day.
What We Evaluate
For BiomniBench-DataAnalysis, we work with domain experts, including first authors of the original papers and industry experts working in the same domain, to curate tasks from published research. Tasks cover different biomedical domains such as oncology, neurodegenerative diseases, and cardiovascular diseases, across different data modalities including transcriptomics, genomics, clinical data, and others. Each task is designed to test both the quality of analytical execution and the depth of biological interpretation.
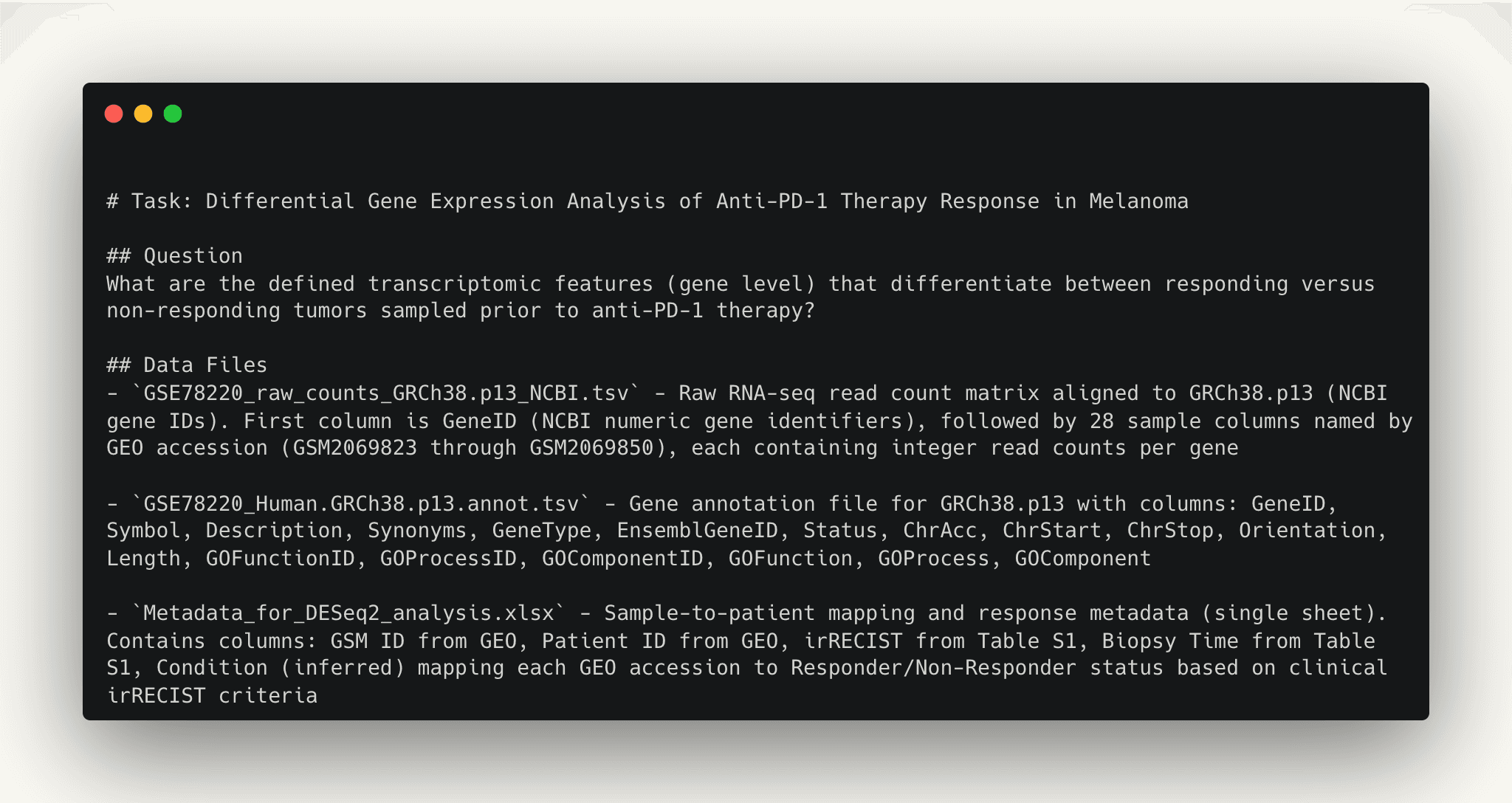
| Dimension | What we're checking | Example criterion |
|---|---|---|
| Data handling | Did the agent load the correct data, apply appropriate filters, and prepare it properly for analysis? | Loads all required files, merges using correct identifiers, filters to the right treatment arm and timepoint, ensures one row per patient, handles measurement scales consistently |
| Tool/method selection | Did the agent choose the right analytical approach for the task? | Chooses Kaplan-Meier with log-rank test for survival analysis; uses Wilcoxon rank-sum for comparing immune populations between survival groups rather than an inappropriate parametric test |
| Statistical rigor | Were tests, thresholds, and corrections applied correctly? | Applies defined significance threshold (p < 0.05), reports exact p-values, applies multiple testing correction (BH-FDR) where appropriate, carries forward interpretation only for statistically significant results |
| Source reliability | Did the agent rely on credible references and validated information? | Provides relevant citations from peer-reviewed literature, databases, and other authoritative sources to support its analytical choices and interpretation |
| Reasoning chain | Was the overall analytical logic coherent, and did the interpretation connect to biology? | Correctly frames the question (e.g., survival-associated vs. predictive biomarkers), avoids causal overclaims, connects findings to biological mechanisms (e.g., CD40-driven antigen presentation), acknowledges study limitations such as lack of longitudinal measurements |
Preliminary Results
BiomniBench-DataAnalysis is still a work in progress. We are sharing preliminary results on a subset of fifteen tasks (Biomni-DA-v0) to provide an early picture of where things stand.
We ran several agents on this subset, alongside two groups of human scientists for comparison. The first group consists of senior to principal-level scientists from large pharmaceutical companies with at least five years of experience. The second group consists of junior scientists with around three years of experience.
Biomni Lab (2026-02-03) scored 65.0, performing comparably to senior pharmaceutical scientists (68.5) and ahead of other agents and the industry scientists group. Edison Analysis scored 52.7, OpenAI Agents SDK (GPT-5.2) scored 51.4, the junior scientists group scored 48.5, and Claude Code (Opus 4.6) scored 47.8.
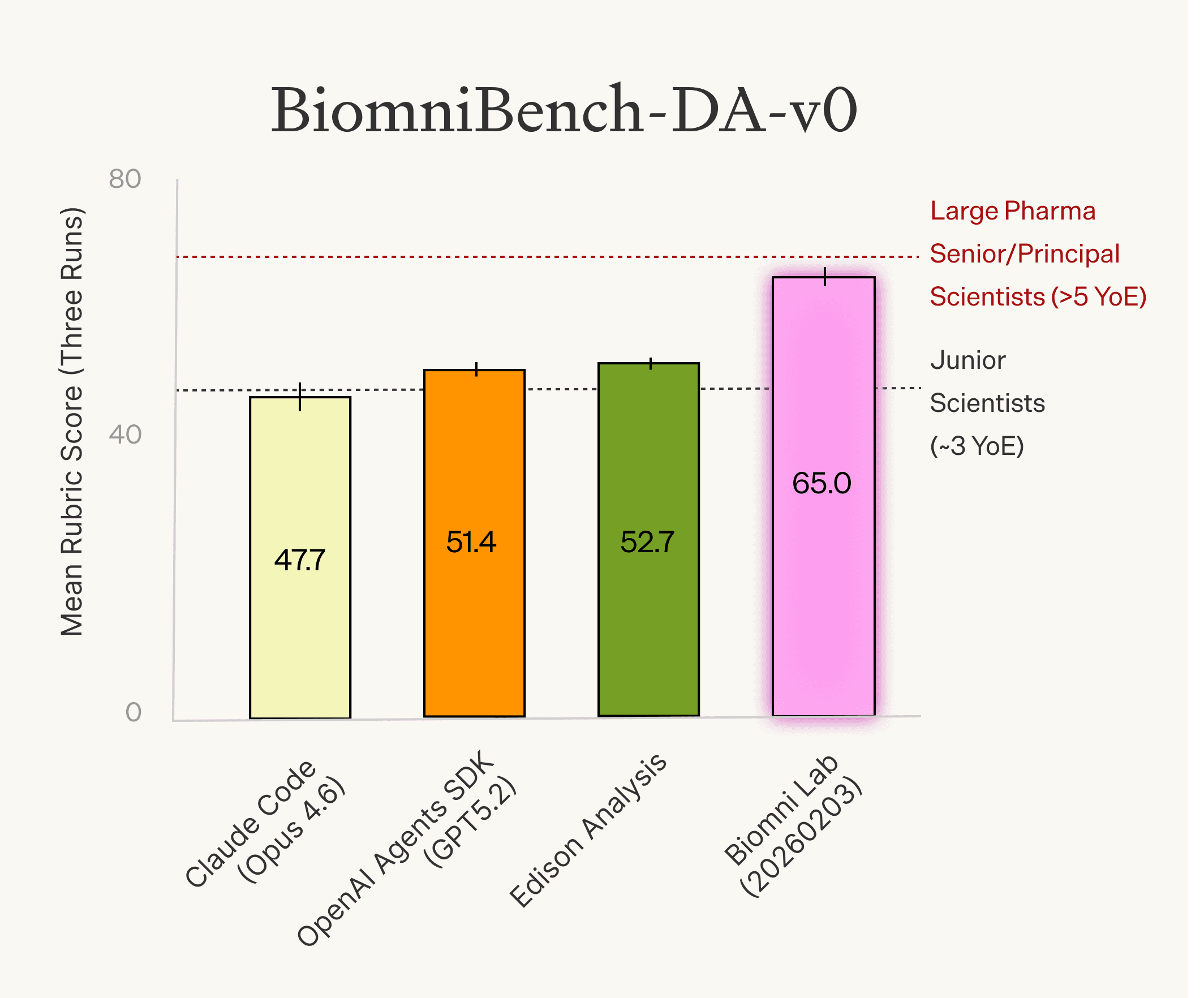
These are early numbers on a small subset, and we expect the picture to evolve as we expand the benchmark.
Our goal with BiomniBench is to build toward an evaluation that reflects the challenges we outlined earlier: covering diverse biological domains and task types from real-world use cases, evaluating multi-step analytical processes rather than just final outputs, and accommodating the reality that many biological questions have more than one valid answer. Our benchmark won't be perfect, but it gives us and the community a way to diagnose where agents fall short and measure real progress as we improve them.
What's Next
We are grateful to our early partners and contributors who have helped shape BiomniBench so far.
We are building this as an open effort because we deeply believe that evaluation for biology should be built by the biology community. The hardest parts of trace-based evaluation are selecting good questions and generating robust rubrics, and every sub-domain of biology has its own standards and judgment calls that only practitioners truly understand. We can’t do it alone.
We are looking for biologists, research institutions, pharmaceutical companies, and agent developers who want to contribute tasks, validate rubrics, or test their systems on trace-based metrics.
If you are interested in contributing or staying updated, reach out at contact@phylobio.com.
We are also hiring.